Reference:
Brown, 1996. Comparative analysis of ribonuclease P RNA using gene sequences from natural microbial populations reveals tertiary structural elements. Proc. Natl. Acad. Sci. U.S.A. 93, 3001-3006. [Medline abstract] Full-text HTML version
I. Phylogenetic analysis
RNase P sequences were amplified from natural population bacterial genomic DNA isolated from greenhouse pond water, near-shore lake and a hot spring outflow. These DNAs were used as templates in PCR amplifications with degenerate oligonucleotide primers complementary to highly conserved sequences, based upon the previously known database of about 50 sequences. Amplified sequences of the expected size (~300 b.p.) were cloned and sequenced. As a result, 52 new unique RNase P RNA sequences were determined, doubling the available sequence database.
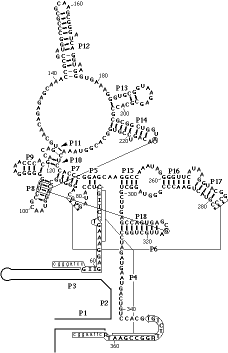
These sequences (available at the RNase P database ) were manually aligned and and analyzed for covariation of nucleotides using the algorithm of Chiu and Kolodziejczak, as described by Gutell, et al. . All of the base pairs shown as heavy bars in the above figure were confimed by this analysis. In addition, tertiary interactions, which were not previously known, were also detected. These are indicated on the figures above and below as lines connecting base pairs in helix P8 with bases in the loops of helices P14 and P18.
Click here <
Base 214 of P14 covaries with the 93/105 base-pair of P8 and base 316 of P18 covaries with the 94/104 base-pair of P8. Both bases 214 and 316 are purines of a conserved GNRA tetraloop motif (Woese, 1990). Each covaries with nucleotides that form Watson-Crick base-pairs in helix P8 of the secondary structure. The covariation is particularly striking in the context of the predicted phylogenetic relationships between the sequences; the A:G/C and G:A/U alternatives are phylogenetically dispersed, i.e. the three bases have changed frequently, as a set, among related sequences. The few exceptions to the correlations (including those that lack either P14 orP18) are phylogenetically clustered and likely represent only a small number of evolutionary events (i.e. they are synaptomorphic).
The straightforward interpretation of these mutual-information correlations is that the covarying nucleotides interact directly to form base-triples, in which A:G/C and G:A/U are acceptable alternatives. Covariation of this type has been observed previously in group I intron sequences (Michel and Westhof, 1990) and shown experimentally to indicate direct interactions (Jaeger et al., 1994).
Click here <
Isosteric structures have been suggested for A:G/C and G:A/U triples in which the loop purine is hydrogen-bonded to the purine of the Watson-Crick base-pair in the minor groove of the helix (Jaeger et al., 1994). Consistent with this hypothesis is the observation that A:G-U sets are present for these bases in some RNase P RNAs, but G:G-U does not occur; the tetraloop purine covaries more strongly with the purine position of the base-pair than with the pyrimidine position. It has been postulated that an additional base-triple might be formed by the adjacent A of the GNRA loop and the purine of the 3'-neighboring base-pair (Michel and Westhof, 1990). In both RNase P and group I intron RNAs, the A of the GNRA loops and the presumptive base-paired partners are extremely conserved, so comparative analysis provides no direct evidence to support the presence of this additional interaction. Nonetheless, in both types of RNA, where the GNRA:base-pair interaction is indicated by phylogenetic covariation, the corresponding adjacent base-pair is conserved and appropriate for base-triple formation with the loop A of GNRA sequences (i.e. A:G/C). Conversely, in RNase P RNAs which lack the ability to form the primary base-triple, the adjacent base-pair in P8 is not conserved as G/C.
II. Tertiary structure
Three-dimensional interpretation. The covariation data for the L14-P8-L18 tertiary interaction were used to model of this domain using the MC-SYM RNA modeling program of Major, et al. The model is generally consistent with available comparative, NMR and crystallographic data for GNRA tetraloops (see Heus and Pardi, 1991, Jaeger et al., 1994, Pley et al., 1994, Cate, et al. 1996). P14 andP18 are aligned coaxially such that their loop nucleotides interact in opposite orientations with the base-paired purines, in the minor groove of P8. <P8, is modeled as proposed for an A:G/C base-triple in group I introns (Jaeger, et al. 1994): the exocyclic A-N6 forms an H-bond with G-N3, and A-N1 pairs with G-N2 <Jaeger et al., 1994) <
The resulting overall structure causes P14 and P18 to form a nearly coaxially stacked continuous
helix, docked at an angle onto P8 <
![]() Back
to intro to RNA structure.
Back
to intro to RNA structure.
![]() This section Still Under Construction
This section Still Under Construction