4.Genomics Informatics or genomic signal processing
Genomic data analysis in the context of post-genome era information processing is an emerging and exciting topic. In the last couple years, high resolution genetic probes have evolved from Human Genome Sequencing Project, which can detect subtle and cryptic genetic aberrations unobtainable with conventional probes, thus, holding great promise for personalized medicine. Among various projects we worked, we have made contributions to the detection of copy number variations (CNVs) first from microarray platform and currently from next generation sequencing (NGS) platform. Recently, we proposed a novel statistical approach that can detect CNVs with better accuracy and more robustness. Our work has been featured in Genetic Engineering & Biotechnology News and Biocompare report.
a. Chen J, Wang YP. A statistical change point model approach for the detection of DNA copy number variations in array CGH data. IEEE/ACM Trans Comput Biol Bioinform. 2009 Oct-Dec;6(4):529-41. PubMed PMID: 19875853; PubMed Central PMCID: PMC4154476.
b. Y.-P.Wang, M. Gunampally, J. Chen, D. Bittel, M. Butler and W.-W. Cai, A Comparison of Fuzzy Clustering Approaches for Quantification of Microarray Gene Expression, Journal of VLSI Signal Processing Special Issue on Machine Learning for Microarray and Sequence Analysis, 50: 305-320, 2008.
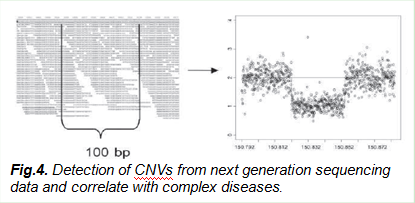
c. Duan J, Zhang JG, Deng HW, Wang YP. CNV-TV: a robust method to discover copy number variation from short sequencing reads. BMC Bioinformatics. 2013 May 2;14:150. PubMed PMID: 23634703; PubMed Central PMCID: PMC3679874.
d. Duan J, Deng HW, Wang YP. Common copy number variation detection from multiple sequenced samples. IEEE Trans Biomed Eng. 2014 Mar;61(3):928-37. PubMed Central PMCID: PMC4165854.
We are currently focusing on two projects: a). Integration of multi-omics data; and b) the analysis of next generation sequencing data. Given the availability of vast amounts of data generated by a variety of high throughput genomic techniques, how to fuse the heterogeneous multimodal genomic information to improve the predication of cancer and the response to therapies is of tremendous significance in biomedicine. We recently received a major NIH R01 award to continue this exciting research in collaboration with Tulane Center for Bioinformatics and Genomics (CBG) on the diagnosis of osteoporosis.

